what are binary expression systems? + bonus problem at the end
GAL4/UAS, Cre-Lox, FLP-FRT, Tet-on/off and more...
Controlling gene expression is a cornerstone of modern biology research. Often, we want to control where a gene is expressed, at what time, or under what conditions. But how do we actually achieve this precise and flexible control?
Binary expression systems are super useful for scientists because they allow very precise control of gene expression and are very scalable and flexible systems.
Binary expression system = two genetic elements (binary) controlling the expression of a gene
Consist of a “driver” and a “responder” element
The driver is a transcription factor that flips the responder element on or off
The driver element is conditionally expressed under a promoter
This promoter could be tissue-specific, or activate upon a certain developmental stage or upon treatment with a drug
The responder element holds the gene we cant to control the expression of
The promoter of the responder element is the “switch” for the driver
In the most simple system, the driver will bind to the promoter and activate transcription, kind of like a simple operon
The expressed gene can be a fluorescent protein, shRNA (knockdown), Cas9 construct, endogenous protein (overexpression) or basically anything!
In diploid model organisms, like Drosophila, C. elegans, or mice, scientists often have driver and promoter strains of the organism that they can cross to get the desired binary system
The binary expression system requires BOTH elements to actually express the gene.
You can mix and match drivers and responders, making these systems extremely flexible. Also, you can have multiple conditions for more specificity.
All of these systems are “leaky” to some extent, because promoters are not perfect. There might be some expression in places you don’t want.
Example:
I want to label the blood vessels of the brain with GFP in a mouse. To do this, I cross a mouse expressing my driver under a sCLDN5 promoter (specific to brain endothelial cells) with a mouse expressing my responder (GFP gene) under a driver response element promoter. The resultant mouse will express the driver only in brain vessels, which will activate the responder only in brain vessels. Thus only brain vessels will have GFP. By the way, if you do this, it might look like this:
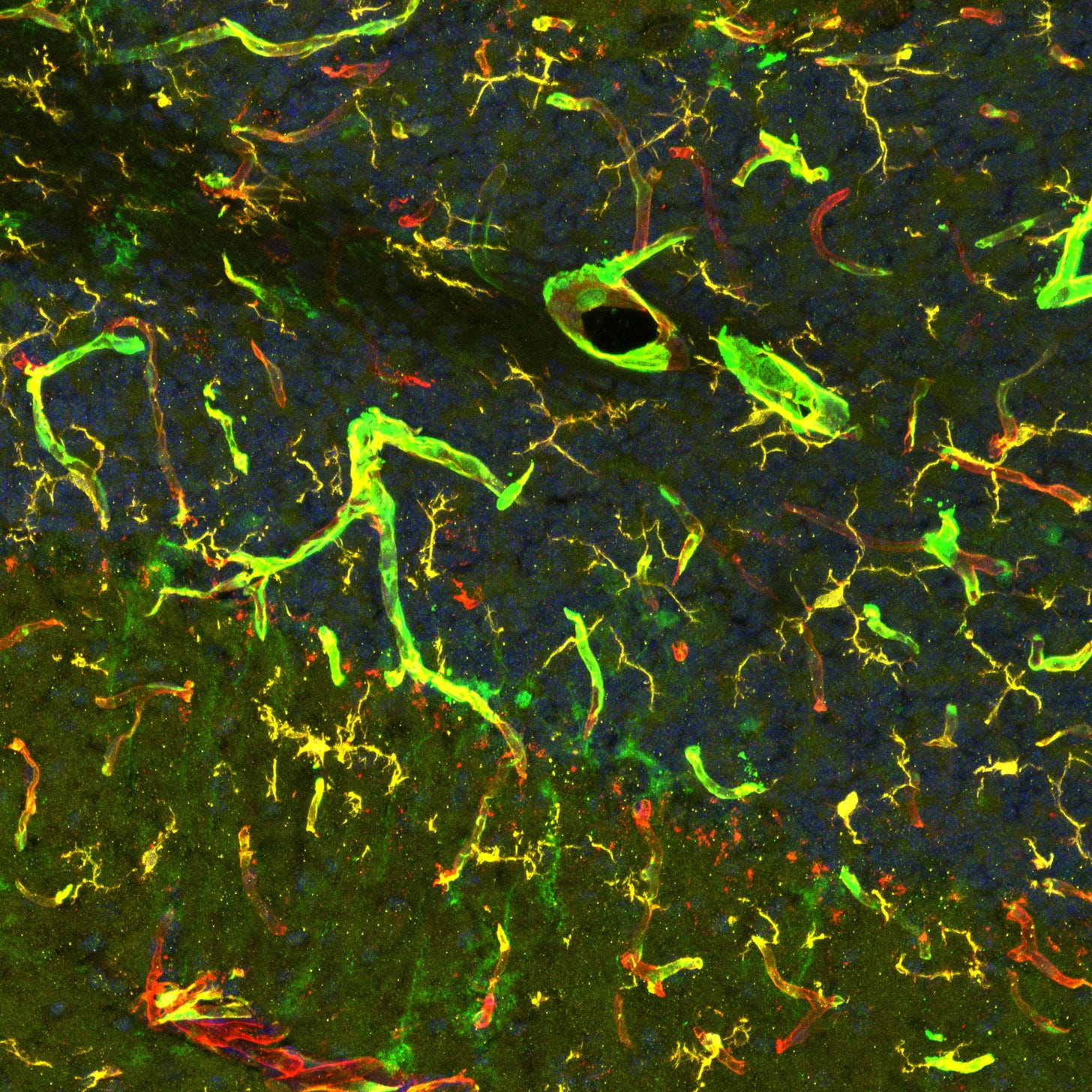
These systems are incredibly diverse. I’ll go over some notable examples, and then a problem at the end.
GAL4/UAS
Probably the simplest system to explain.
GAL4 = driver
UAS = responder
GAL4 binds to the UAS promoter and activates transcription
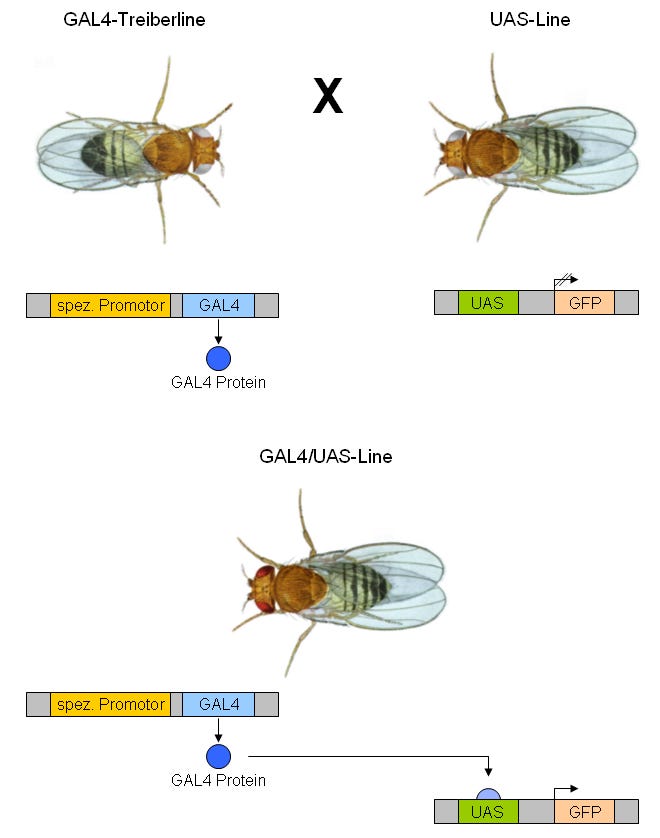
There are many “riffs” or modifications of this system that can be used in combination or by themselves, such as
temperature-sensitive mutants of the GAL4 protein inactivate upon heating
e.g. want to express GFP in brain, but only until the larval stage
GAL80 is an inhibitor of GAL4, so you can get even more specific. —> need this and need NOT that
want to express GFP in all neurons (express ELAV) EXCEPT dopaminergic neurons (express TH) —> ELAV-GAL4 TH-GAL-80/UAS-GFP
some versions of GAL4 can be switched on and off with drugs
e.g. want to express GFP in the brain only after adulthood
because Gal4, like many TFs, is a dimer, you can make “split” Gal4. This means you need two separate halves to join together to work. You can put both halves under different promoters. —> need this AND that.
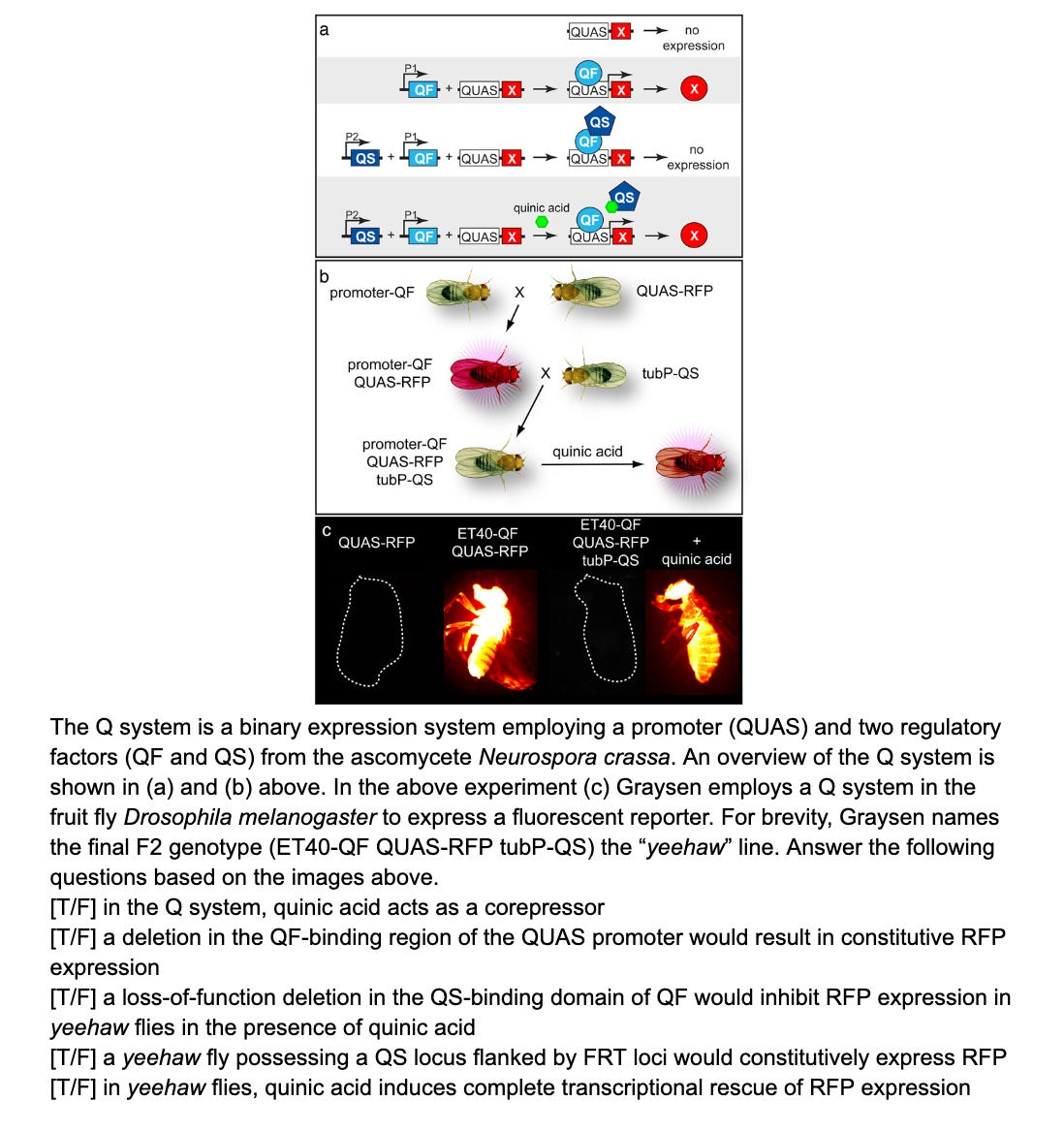
Other systems like Q system and LexA-LexAop are similar in structure one you get the concept
Tetracycline-on/off
Comes from an antibiotic response element in bacteria. When tetracycline or doxycycline is added, the driver protein is either inactivated (tet-off) or activated (tet-on).
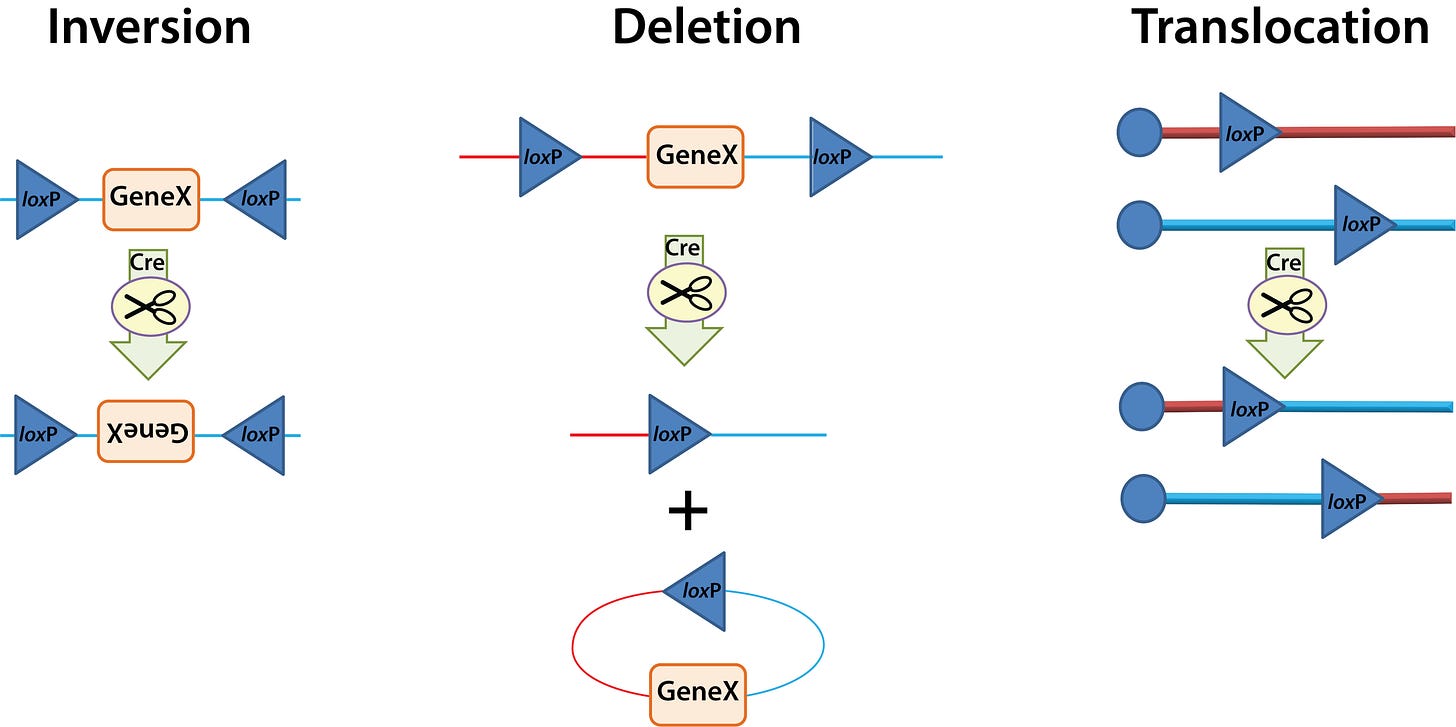
Cre-Lox and FLP-FRT
Recombination based systems are not strictly binary expression systems, but they work in the same way. In recombination systems, instead of a promoter, the responder element contains cut sites for a recombinase, turning the gene on or off by cutting and rearranging the DNA. For example,
Driver = Cre recombinase
Responder = gene of interest (GOI) flanked by loxP “cut” sites
Response = when Cre is expressed, it cuts at the LoxP sites, turning the GOI on/off via any of several mechanisms, shown below.
The “floxed” element (flanked by loxP cut sites) can be the gene itself, a stop codon, a promoter or enhancer, intron, etc.
FLP-FRT is basically the same, just with different recombinase and cut sites.
This was a very brief overview of binary expression systems. They’re extremely useful for many research applications and can be implemented in to basically any model organism. If you have any questions, ask in the comments! I’ve been able to work with many of these systems before, and they are extremely useful.
Quiz time!
Put your answers or questions in the comments and stay tuned!